Readme from the project
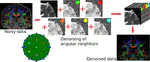
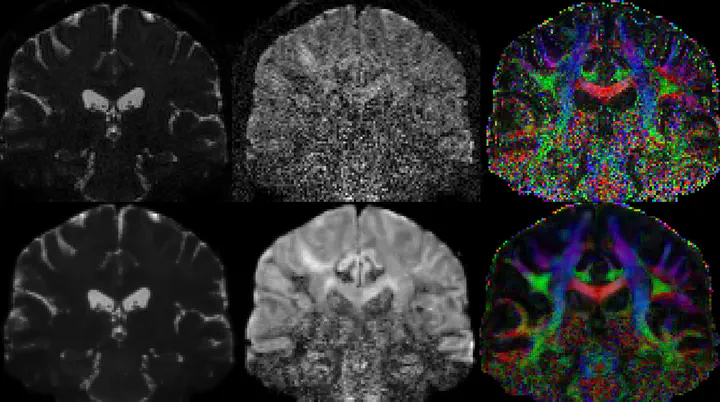
Non Local Spatial and Angular Matching (NLSAM) denoising
The reference implementation for the Non Local Spatial and Angular Matching (NLSAM) denoising algorithm for diffusion MRI.
Quick links
You can find the latest documentation and installation instructions over here with a downloadable version of the documentation here.
How to install
If you have a working python setup already, the next command should give you everything you need.
pip install nlsam
There are also Dockerfiles in the folder, for which you’ll need to install docker if you want to build upon it for pipeline processing.
You can also download the datasets used in the paper over here.
Using the NLSAM algorithm
The process is to first transform your data to Gaussian distributed signals if your dataset is Rician or Noncentral chi distributed and then proceed to the NLSAM denoising part itself.
A quickstart example call would be
nlsam_denoising dwi.nii.gz dwi_nlsam.nii.gz 1 bvals bvecs 5 -m mask.nii.gz
For more fine grained control and explanation of arguments, have a look at the possible command line options with nlsam_denoising –help
You can find a detailed usage example and assorted dataset to try out in the example folder.
Questions / Need help / Think this is great software?
If you need help or would like more information, don’t hesitate to drop me a line at samuel.st_jean@university, where university needs to be replaced with med.lu.se
References
The NLSAM denoising algorithm itself is detailed in
St-Jean, S., Coupé, P., & Descoteaux, M. (2016). “Non Local Spatial and Angular Matching : Enabling higher spatial resolution diffusion MRI datasets through adaptive denoising” Medical Image Analysis, 32(2016), 115–130. DOI URL
The bias correction framework is a reimplementation of
Koay, CG, Özarslan, E and Basser, PJ A signal transformational framework for breaking the noise floor and its applications in MRI, Journal of Magnetic Resonance, Volume 197, Issue 2, 2009
The automatic estimation of the noise distribution is computed with
St-Jean S, De Luca A, Tax C.M.W., Viergever M.A, Leemans A. (2020) “Automated characterization of noise distributions in diffusion MRI data.” Medical Image Analysis, October 2020:101758. doi:10.1016/j.media.2020.101758
And here is a premade bibtex entry.
@article{St-Jean2016a,
author = {St-Jean, Samuel and Coup{\'{e}}, Pierrick and Descoteaux, Maxime},
doi = {10.1016/j.media.2016.02.010},
journal = {Medical Image Analysis},
pages = {115--130},
title = {{Non Local Spatial and Angular Matching : Enabling higher spatial resolution diffusion MRI datasets through adaptive denoising}},
volume = {32},
year = {2016}
}
